How pH Affects Separation of Oligonucleotides in Ion Exchange Chromatography
Matthew Crawford
Purifying oligonucleotides for research, diagnostics, or therapeutic applications often relies on ion exchange chromatography (IEX) due to its highly biocompatible properties during separation and good resolution. Among the various chromatographic methods available, IEX is particularly powerful because it separates molecules based on charge, offering both sensitivity and scalability from analytical research to production. To fully understand why oligonucleotides behave the way they do during ion exchange and how that affects separation/purification method development, we need to explore two critical aspects of IEX: the role of pH in elution and (relatedly) the impact of charge on separation.
The Role of pH in Ion Exchange Chromatography
In ion exchange chromatography, separation occurs because biomolecules interact electrostatically with a charged stationary phase. For oligonucleotides, the phosphate backbone carries a negative charge under most conditions. This charge allows oligos to bind strongly to anion exchange resins, which are positively charged. pH plays a crucial role because it influences both:
The charge state of the oligonucleotide:
- Each phosphate group in DNA and RNA has a pKa of around 1.0–1.5. This means that at nearly all biologically relevant pH values (above ~2), the backbone is fully deprotonated and negatively charged.
- As a result, changes in pH don’t significantly alter the net negative charge of the oligo itself, but they can affect other factors (like end modifications, protecting groups, or impurities) that may shift separation profiles.
The charge of the resin functional groups:
- Strong anion exchange resins typically use quaternary amines (Q-resins), which remain positively charged across a broad pH range.
- At lower pH, weaker anion exchangers (like DEAE resins) may lose positive charge, reducing their binding capacity and affecting resolution of bound compounds.
While the oligo’s charge is relatively stable, the resin chemistry and buffer environment determine how tightly it binds and how easily it elutes, affecting the separation profile.
How Charge Affects Separation
Since oligonucleotides carry multiple negative charges, their retention on an anion exchange column is strong and proportional to the number of phosphate groups, which is tied to the sequence length. Longer oligonucleotides (or those with fewer modifications) often bind more tightly. Charge differences can also distinguish between:
- Full-length oligonucleotides vs. truncated species. Truncated products have fewer phosphates (and therefore less charge), leading to weaker binding and earlier elution.
- This is especially relevant to the purification of synthetically made oligonucleotides.
- Impurities or chemically modified species. Variations in backbone chemistry, capping groups, or conjugations can shift the effective charge density and alter retention.
- This is especially relevant to conjugates that contain oligonucleotides, such as antibody-oligonucleotide conjugates or peptide-oligonucleotide conjugates.
During elution, a salt gradient (often NaCl, KCl, or NaClO4) competes with the negatively charged oligos for binding to the resin. Higher salt concentrations displace stronger binders, creating a separation based on charge density.
Why Anion Exchange is the Method of Choice for Oligonucleotides
Because the phosphate backbone of DNA and RNA is inherently negatively charged at nearly all working pH values, anion exchange chromatography is the natural choice for separation. Cation exchange, by contrast, would require the oligo to carry a net positive charge, which only occurs under extreme and impractical conditions (very low pH, where the backbone loses deprotonation) where the oligonucleotides would be susceptible to degradation via various chemical routes. Anion exchange offers:
- High resolution between oligos differing by just a single nucleotide.
- Robust binding under a wide pH range.
- Scalability for both analytical and preparative purification.

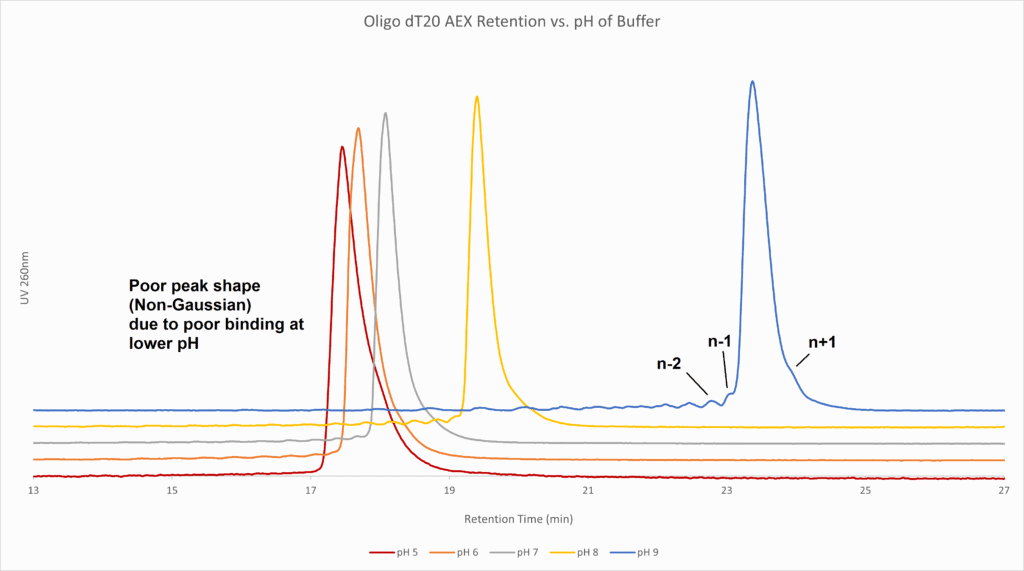
Figure 1: Anion exchange chromatography on synthetic dT20 (DNA poly-T 20-mer) at various pH values from 0 to 1M NaCl. Note the improved retention of the target oligo as well as improved resolution of n-1/n+1 sequence truncations. This pH screening would be followed by NaCl gradient focusing at pH 9.0 to further separate the contaminants from synthesis.
Takeaways
pH influences ion exchange chromatography of oligonucleotides primarily by tuning the resin environment and buffer conditions, rather than altering the fundamental charge state of the oligonucleotide backbone. Since oligos are strongly negatively charged under nearly all practical conditions, they bind efficiently to positively charged anion exchange resins. Differences in charge density—arising from length, truncation, or modifications—allow for precise separation, with salt gradients enabling controlled elution.
That’s why anion exchange chromatography remains the gold standard for purifying oligonucleotides. At Rilas, we leverage the natural chemistry of nucleic acids for reliable, high-resolution separation of your analytes for biocompatible downstream experimentation.
